SCN Transcriptomic Meta-Analysis
Interactive Data Visualization
Data and Jupyter/IPython notebooks to examine enriched transcripts in the SCN via Meta-analysis.
The suprachiasmatic nuclei (SCN) in the brain represents the master circadian pacemaker in mammals. It controls the timing of many aspects of physiology and behaviour,
as well as coordinating other peripheral clocks in organs throughout the body.
The expression of genes in SCN has often focused on the identification of rhythmically expressed genes.
A different approach is taken here: to look at those selectively enriched in the SCN, by comparing the transcriptome of the SCN to that of the brain in general.
This will help to find genes that play key roles in SCN physiology or may refine SCN-specific drivers.
ORIGINAL REFERENCE: Laurence A. Brown, John Williams, Lewis Taylor, Ross J. Thomson, Patrick M. Nolan, Russell G. Foster, Stuart N. Peirson; Meta-analysis of transcriptomic datasets identifies genes enriched in the mammalian circadian pacemaker, Nucleic Acids Research,
Volume 45, Issue 17, 29 September 2017, Pages 9860–9873, https://doi.org/10.1093/nar/gkx714
All the code to run the original analysis as found in the NAR paper is included with the paper,
as are the data.
- Versions of the analysis will be found on GitHub
79
Microarray files used (4 platforms)
17
RNA-Seq files
31, 684
MGI gene symbols with data
426 genes
Effect Sizes over 2, (q<0.01)
Continuous Open Mouse Phenotyping of Activity and Sleep Status (COMPASS)
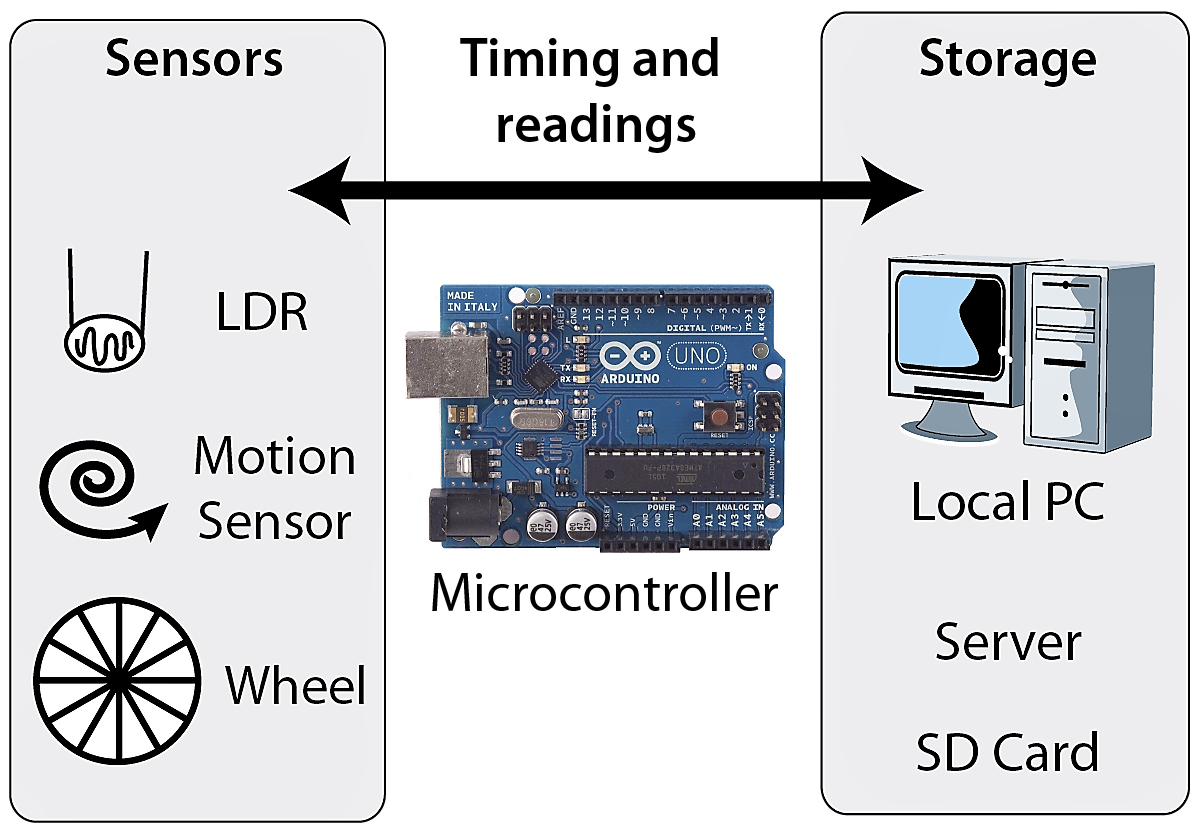
An open and low-cost option for monitoring actigraphy and estimating sleep in mice
This project represents our attempt to provide an affordable and non-invasive
method to assess activity and (by immobility) sleep in laboratory mice. The publication in Wellcome Open Research describes some
of the many options for data collection and analysis.
ORIGINAL REFERENCE: Brown LA, Hasan S, Foster RG and Peirson SN.
COMPASS: Continuous Open Mouse Phenotyping of Activity and Sleep Status [version 2; referees: 4 approved].
Wellcome Open Res 2017, 1:2 (doi: 10.12688/wellcomeopenres.9892.2)
https://wellcomeopenresearch.org/articles/1-2/v2
- Updated versions of the analysis will be found on GitHub
- Further details on constructing a COMPASS system can be found HERE (in progress)
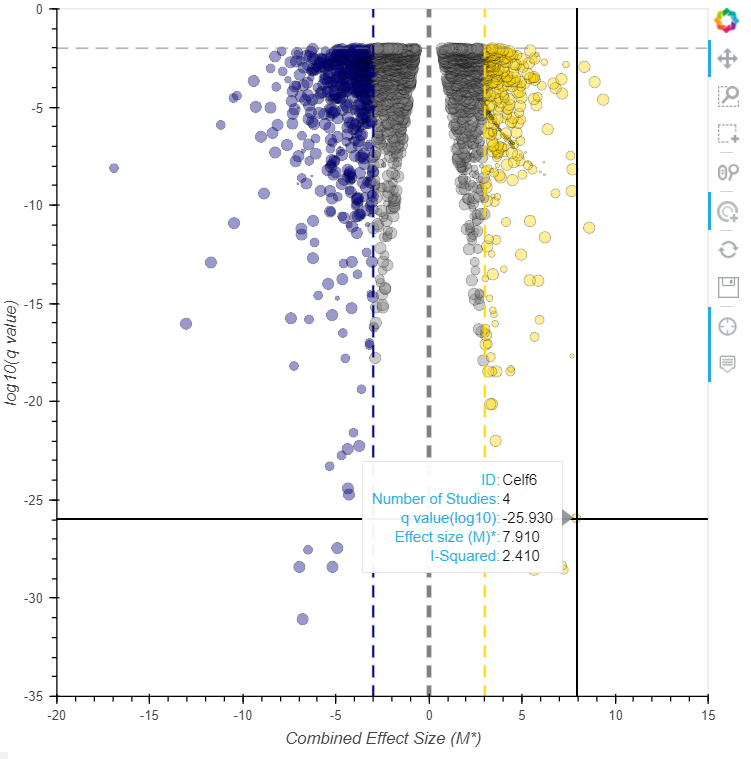 Click to access current app.
(using modern browser, Firefox, Chrome or Edge should be fine)
Click to access current app.
(using modern browser, Firefox, Chrome or Edge should be fine)

